 Thirdly, Java requires you to allocate memory when you start ImageJ. If you download ImageJ from the ImageJ or Fiji websites, you can specify which operating system you have and it will download the correct java along with it. Secondly, even if you have a 64-bit machine, your computer may or may not have 64-bit Java installed. 64-bit systems do not have this limitation and are therefore limited by the amount of RAM the computer has. If nothing is shown, the system is likely 32-bit. If you don't know whether your computer is 32-bit or not, right click on My Computer and go to the Properties menu and look for 32-bit or 64-bit somewhere on the resulting page. Firstly, Java limits 32-bit computers to around 1.5 GB of memory. A couple of important points about memory. ImageJ is written in the Java programming language, so limitations in Java apply to ImageJ. This frustrates some users, but I feel that it is better to understand the memory limitations of a particular task rather than lock up the computer at some unknown point during analysis. ImageJ manages this process in a relatively transparent fashion. I'll probably always use Fiji in future as it's just easier than downloading all the plugins myself.Whether you like it or not, managing computer memory is an important part of all scientific image processing tasks. The 3D viewer seems to work better on this version too. You can manually set an inter-slice distance though and this seems to sort it. It does however still have the problem when importing image files that it doesn't know the inter-slice distance (it is in the metafile and other software finds it! grr). It features a segmentation editor which is brilliant for generating a separate labels file for rendering in 3D. Fijiįiji (Fiji Is Just ImageJ¶ - Batteries included) is ImageJ packaged along with Java 3D and a load of plugins pre-organised into menus for ease of use. "Amygdala and hippocampal volumes in adolescents and adults with bipolar disorder". Kaufman, J Martin, A Whiteman, R Zhang, JH Gore, JC Charney, DS Krystal, JH et al. This interpolates the volume between slices which should be more accurate than assuming it's a perfect geometric shape.This method runs a pilot first then the actual determination of volume using the pilot as an estimate to give a more accurate value.There are, however, two benfits to this plugin over stock ImageJ: You then go through slice by slice drawing around the regions of interest in a similar manner to the one above. the amygdala in an adult human is on average 2117☖0 mm³ ‖). For this estimate literature values are always good (e.g. This plugin starts by asking you how many slices you can see the object on and an estimate of the volume. Volumest is a plugin for ImageJ which stands for VOLUMe ESTimation. The width of this volume is the slice thickness. §In MRI a 2D image (or slice) acquires signal from a volume and presents it as a 2D image. 
‡You need to set your scale first otherwise the area will be in pixels not cm³. aren't ordinary geometric shapes this is not necessarily the most accurate method. Using the ROI manager you can then store this information for each slice individually and then use the measure function to find the area of your ROI on each slice.‡ Multiplying this number by the thickness of your slices gives an estimate of the volume you are measuring.§ Since organs etc. By holding Shift you can create a region of interest composed of multiple parts. Using stock ImageJ with no plugins you can create a region of interest (ROI) using the drawing tools.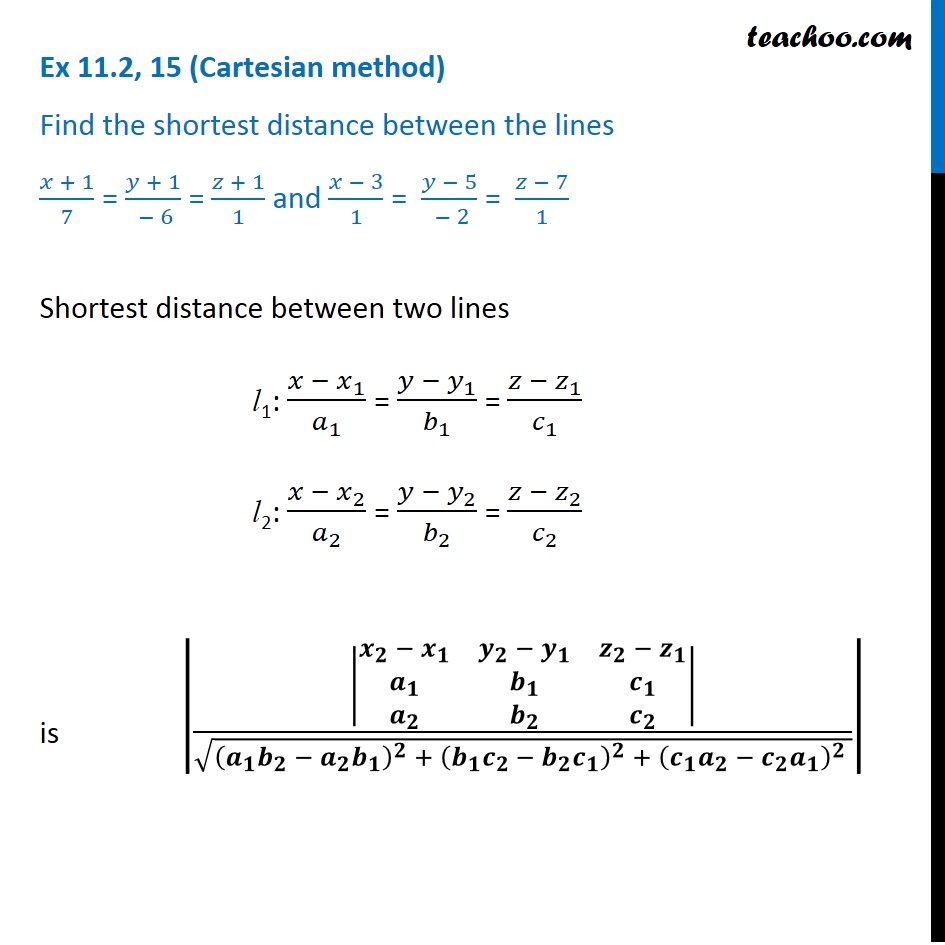 
This is very annoying when it renders your 3D volume as almost flat :( ImageJ is not very good at getting the correct inter-slice distance from your original image files. A stack is a series of 2D images stacked on top of each other to make a 3D image. ImageJ deals with volumes by storing them as stacks. ImageJ is a free, java based, image analysis package provided by the NIH.† As it's Java based it's platform independent as long as you have an up-to-date Java runtime. 
*There are differences between these two methods of acquisition but that would require a whole blog post on it's own. Here are my experiences of trying to measure volumes on MRI images. After all when someone has brain cancer - "How big is the tumour?" can be a very important question. As far as this blog post is concerned these are the same.*Ī common measure in clinical MRI is the volume of an organ or part of an organ. In which I describe the complexities of analysing 3D images.ģD image analysis just seems to be one of those things that is harder to do than I'd like it to be.īefore starting it is worth pointing out that with MRI you can acquire volumes in two ways - traditional 2D with multiple slices or a 3D scan. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
